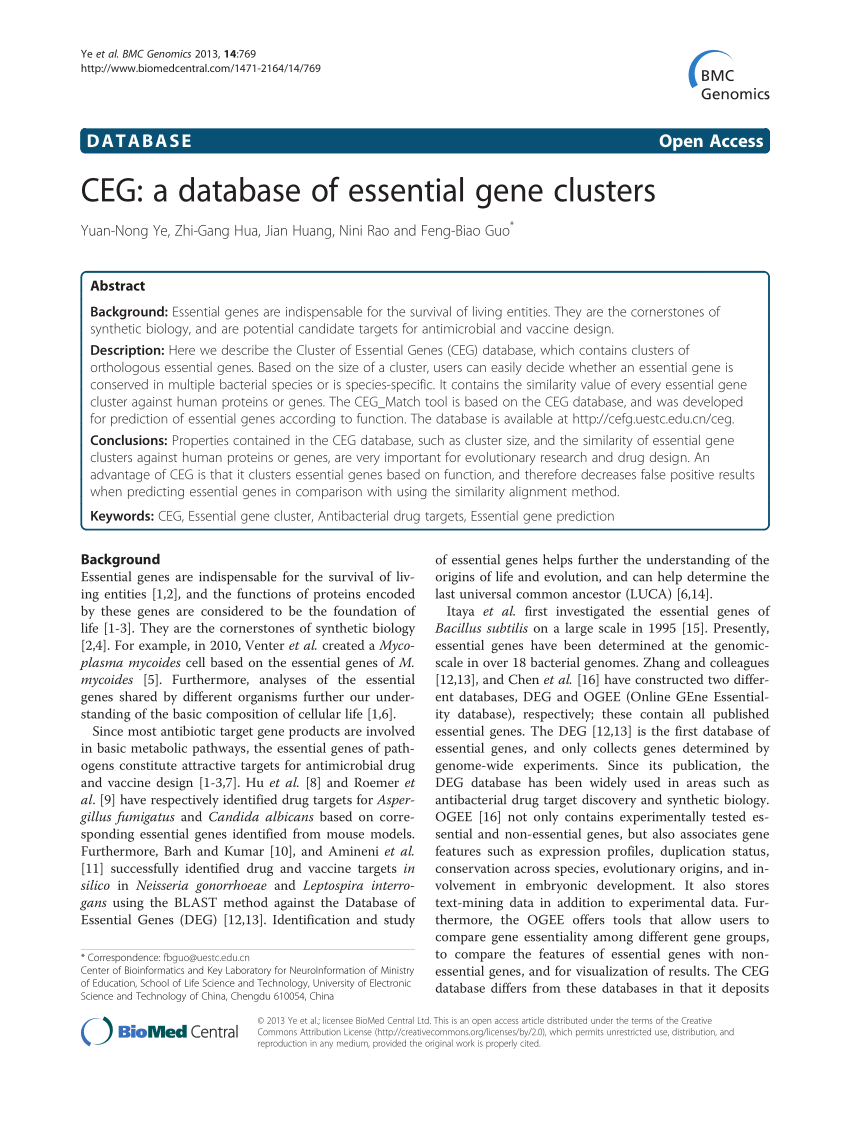
![PDF] DEG 10, an update of the database of essential genes that includes both protein-coding genes and noncoding genomic elements | Semantic Scholar PDF] DEG 10, an update of the database of essential genes that includes both protein-coding genes and noncoding genomic elements | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/f931271dc718f971bfb7b77cdc9d7481929c0392/3-Table1-1.png)
PDF] DEG 10, an update of the database of essential genes that includes both protein-coding genes and noncoding genomic elements | Semantic Scholar

Human and mouse essentiality screens as a resource for disease gene discovery | Nature Communications
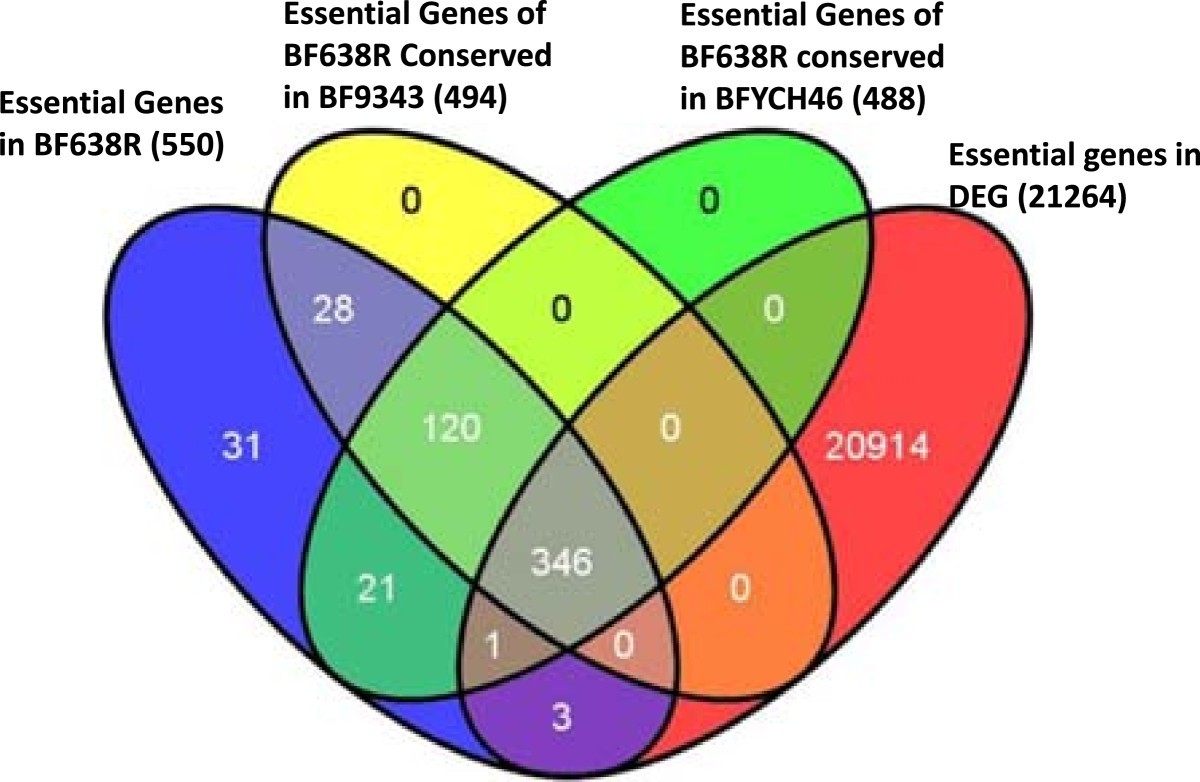
Identification of genes required for the survival of B. fragilis using massive parallel sequencing of a saturated transposon mutant library | BMC Genomics | Full Text
Accurate prediction of human essential genes using only nucleotide composition and association information
GitHub - mpi2/essential-genes-database: Data base load scripts to support the IMPC Essential Genes portal
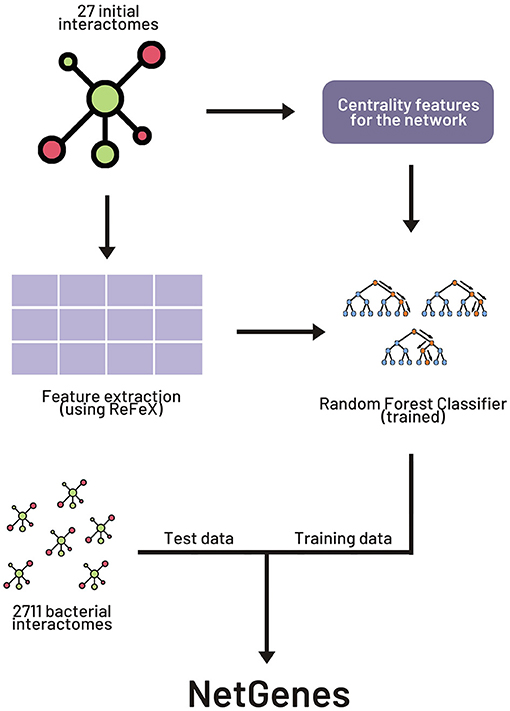
Frontiers | NetGenes: A Database of Essential Genes Predicted Using Features From Interaction Networks
![PDF] DEG 15, an update of the Database of Essential Genes that includes built-in analysis tools | Semantic Scholar PDF] DEG 15, an update of the Database of Essential Genes that includes built-in analysis tools | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/3ac24e345447677a5aa40a449ccfaac7d8b46dfd/2-Figure1-1.png)
PDF] DEG 15, an update of the Database of Essential Genes that includes built-in analysis tools | Semantic Scholar
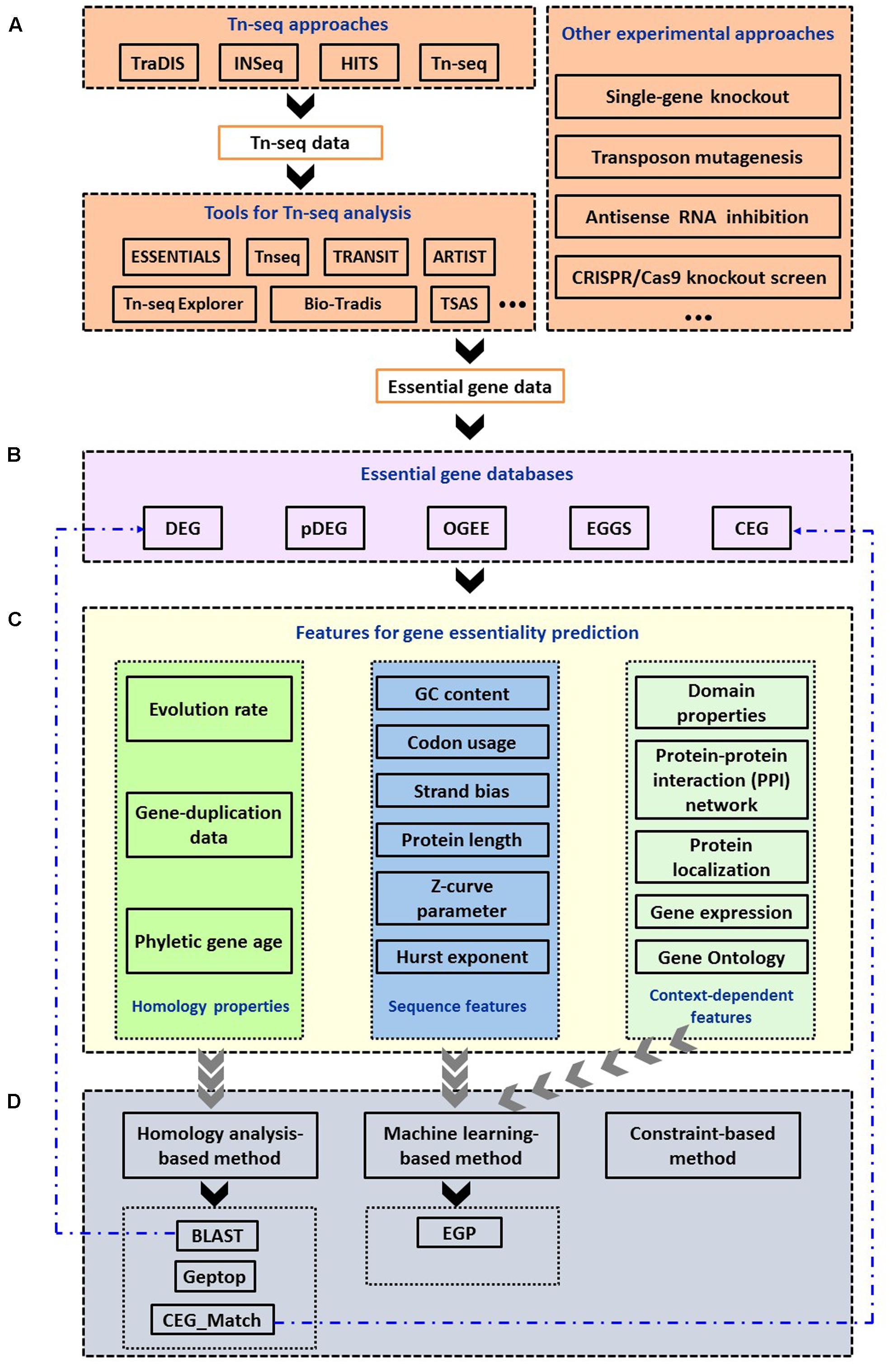
Frontiers | A Comprehensive Overview of Online Resources to Identify and Predict Bacterial Essential Genes
In Silico Insights into the Symbiotic Nitrogen Fixation in Sinorhizobium meliloti via Metabolic Reconstruction | PLOS ONE
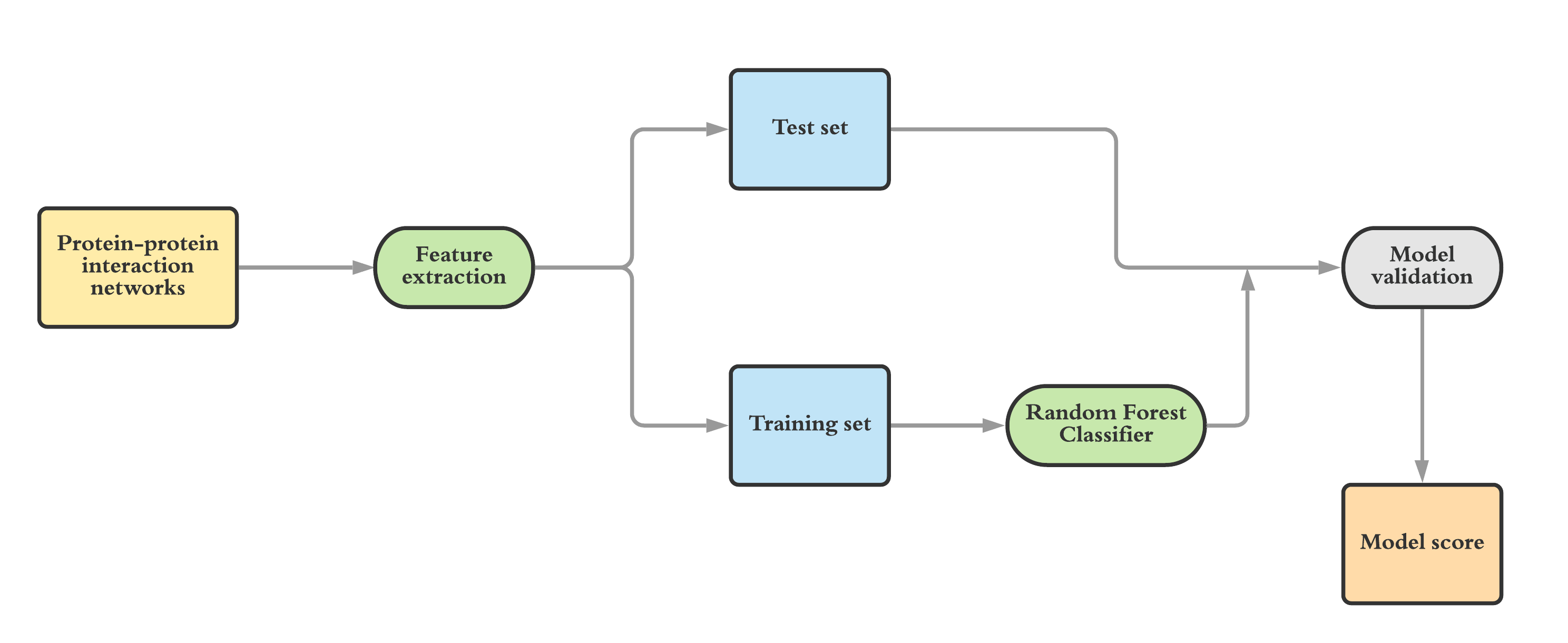
Predicting Essential Genes through Network Approach: Deciphering basis of Life | Robert Bosch Center for Data Science and Artificial Intelligence
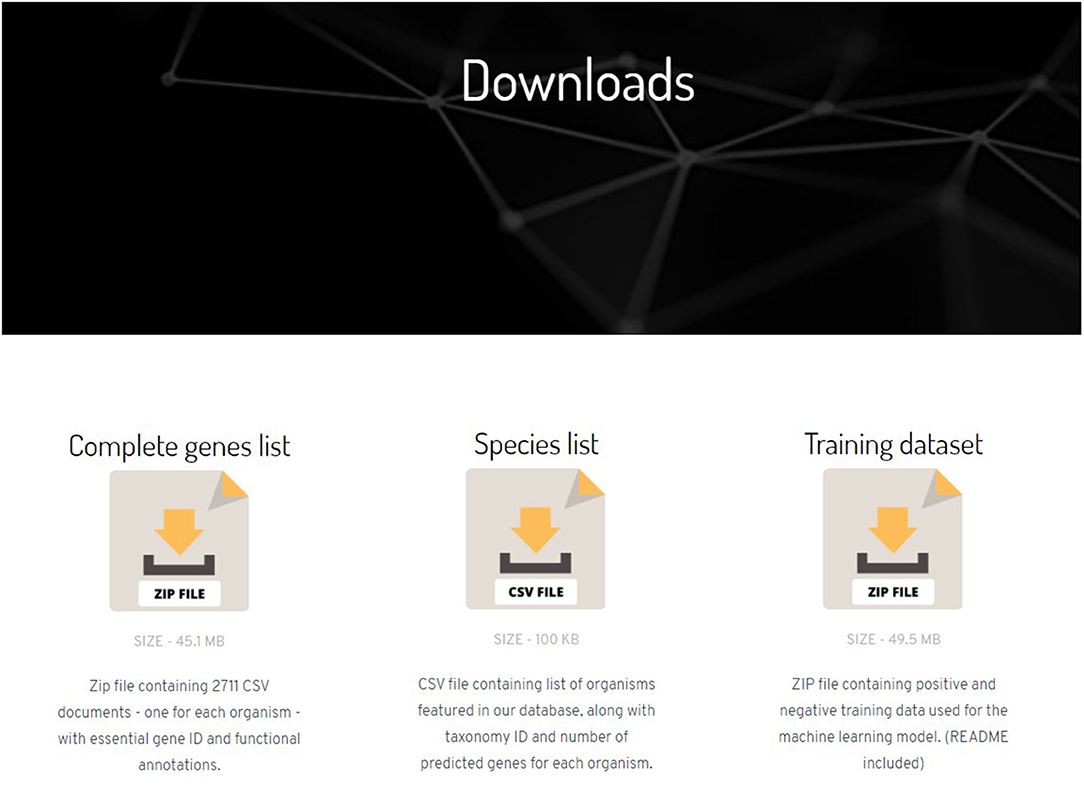
Frontiers | NetGenes: A Database of Essential Genes Predicted Using Features From Interaction Networks

![PDF] DEG 5.0, a database of essential genes in both prokaryotes and eukaryotes | Semantic Scholar PDF] DEG 5.0, a database of essential genes in both prokaryotes and eukaryotes | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/2dd7b6cfcffd83ce52695b2b984e7223517486f5/2-Table1-1.png)

![PDF] Network-based methods for predicting essential genes or proteins: a survey | Semantic Scholar PDF] Network-based methods for predicting essential genes or proteins: a survey | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6fe668c9e79cb7dc3485ce4954558e7e676926ac/3-Table1-1.png)






